Embeds the data set with the specified (relative) term contribution
Source:R/add-functions.R
add_term.RdAdds the contribution of a specific term to the
linear predictor to the data specified by newdata.
Essentially a wrapper to predict.gam, with type="terms".
Thus most arguments and their documentation below is from predict.gam.
add_term(newdata, object, term, reference = NULL, ci = TRUE, se_mult = 2, ...)Arguments
- newdata
A data frame or list containing the values of the model covariates at which predictions are required. If this is not provided then predictions corresponding to the original data are returned. If
newdatais provided then it should contain all the variables needed for prediction: a warning is generated if not. See details for use withlink{linear.functional.terms}.- object
a fitted
gamobject as produced bygam().- term
A character (vector) or regular expression indicating for which term(s) information should be extracted and added to data set.
- reference
A data frame with number of rows equal to
nrow(newdata)or one, or a named list with (partial) covariate specifications. See examples.- ci
logical. Indicates if confidence intervals should be calculated. Defaults toTRUE.- se_mult
The factor by which standard errors are multiplied to form confidence intervals.
- ...
Further arguments passed to
predict.gam
Examples
library(ggplot2)
ped <- as_ped(tumor, Surv(days, status)~ age, cut = seq(0, 2000, by = 100))
pam <- mgcv::gam(ped_status ~ s(tend) + s(age), family = poisson(),
offset = offset, data = ped)
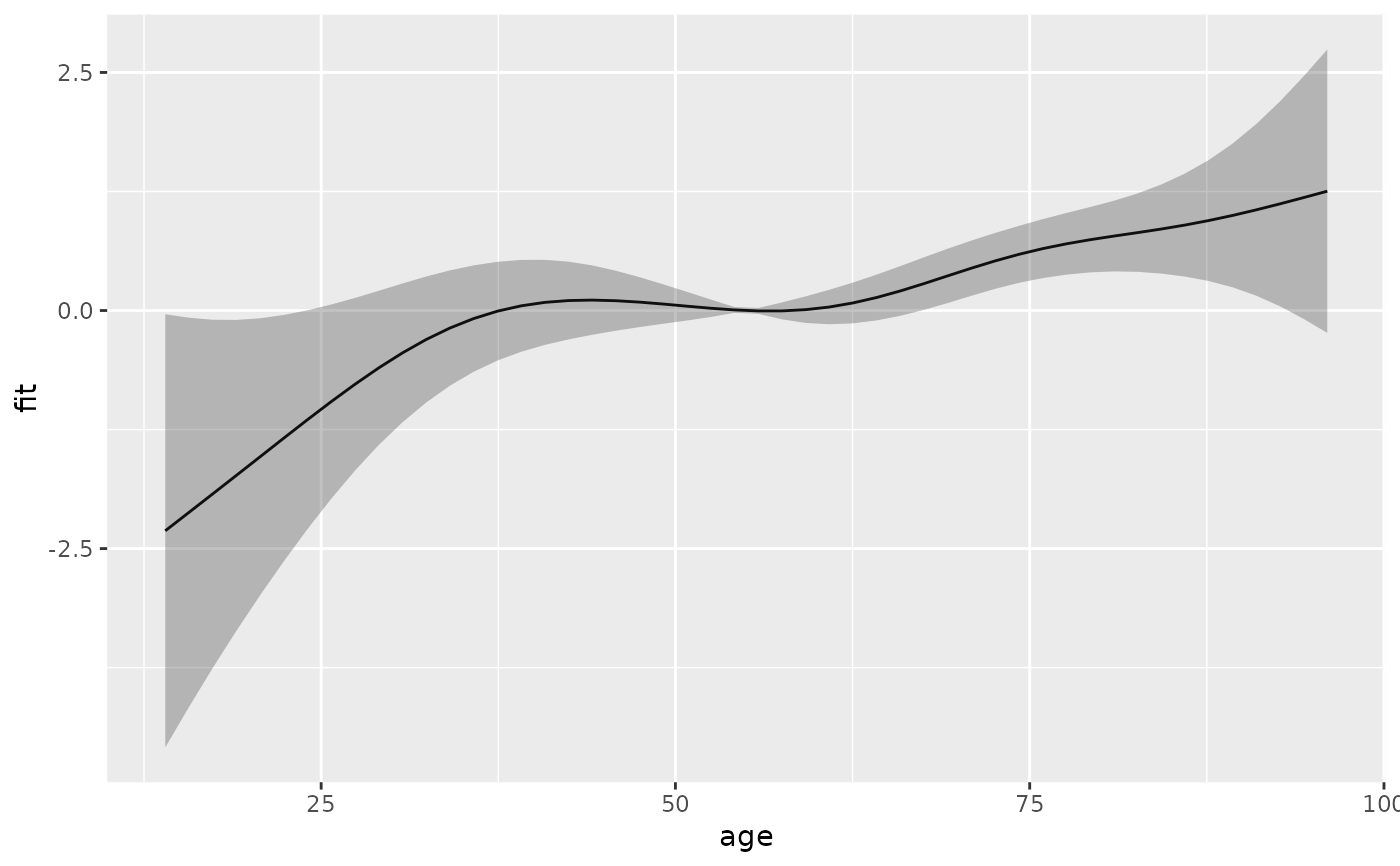
#term contribution for sequence of ages
s_age <- ped %>% make_newdata(age = seq_range(age, 50)) %>%
add_term(pam, term = "age")
ggplot(s_age, aes(x = age, y = fit)) + geom_line() +
geom_ribbon(aes(ymin = ci_lower, ymax = ci_upper), alpha = .3)
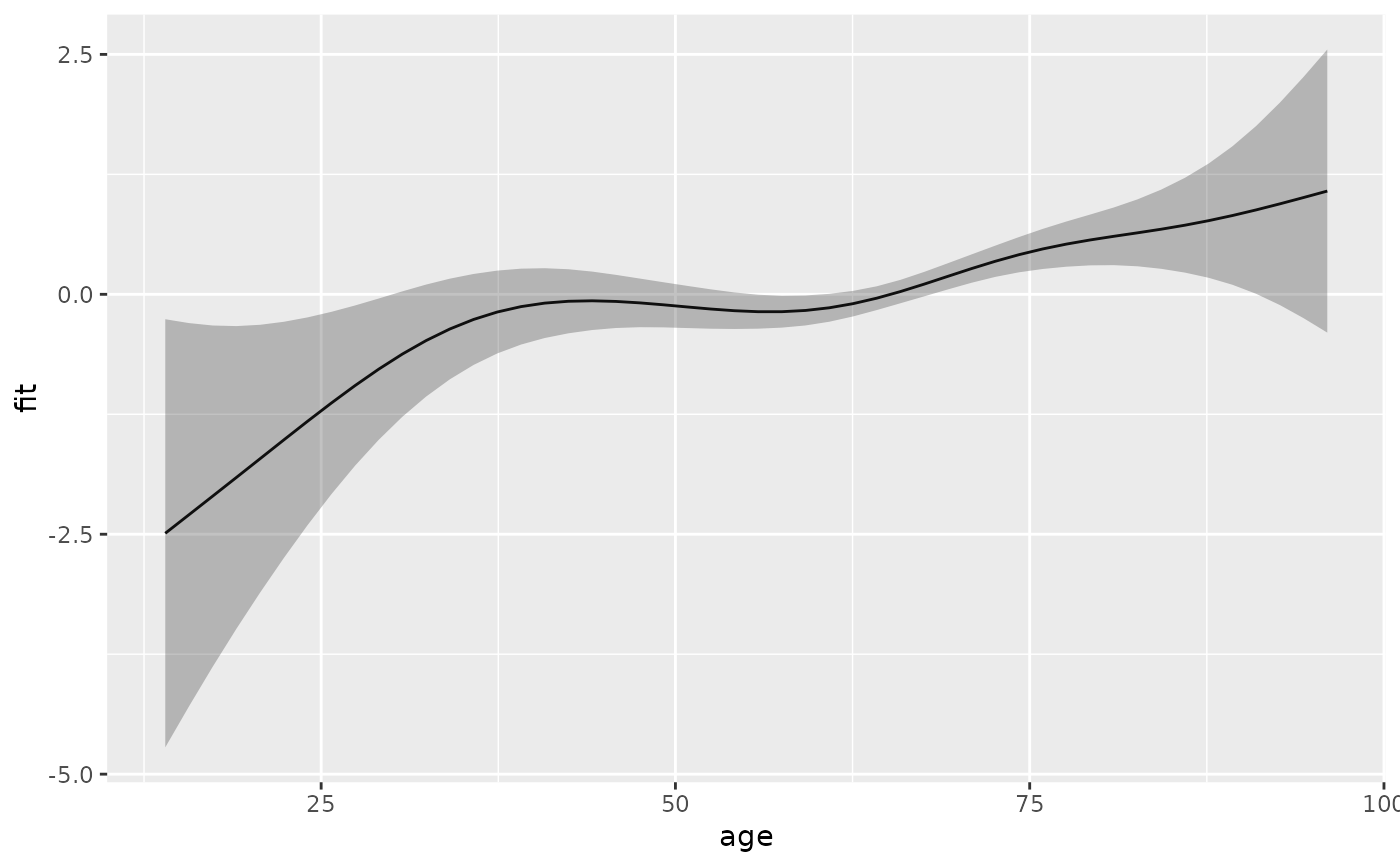 # term contribution relative to mean age
s_age2 <- ped %>% make_newdata(age = seq_range(age, 50)) %>%
add_term(pam, term = "age", reference = list(age = mean(.$age)))
ggplot(s_age2, aes(x = age, y = fit)) + geom_line() +
geom_ribbon(aes(ymin = ci_lower, ymax = ci_upper), alpha = .3)
# term contribution relative to mean age
s_age2 <- ped %>% make_newdata(age = seq_range(age, 50)) %>%
add_term(pam, term = "age", reference = list(age = mean(.$age)))
ggplot(s_age2, aes(x = age, y = fit)) + geom_line() +
geom_ribbon(aes(ymin = ci_lower, ymax = ci_upper), alpha = .3)